Difference between revisions of "Repository of scripts used by IRIDIA members"
From IridiaWiki
Jump to navigationJump to search| Line 14: | Line 14: | ||
| |
| |
||
|-+ src (contains the sources of the algorithms) |
|-+ src (contains the sources of the algorithms) |
||
| + | | |
||
| + | |-+ analysis (contains the R script to boxplot the experimental data) |
||
</pre> |
</pre> |
||
| Line 33: | Line 35: | ||
tai100a 21059006 |
tai100a 21059006 |
||
tai100b 1185996137 |
tai100b 1185996137 |
||
| ⚫ | |||
| + | |||
| + | |||
| + | in the files "algorithm.txt" I put the algorithms details |
||
| ⚫ | |||
| + | landau:~/Desktop/Parallel-QAP-2cpu/out/analysis/2-opt mmanfrin$ head algorithms.txt |
||
| ⚫ | |||
| + | PIR-2opt PIR PIR 2 Par 2 |
||
| ⚫ | |||
| + | SEQ-2opt SEQ SEQ 2 Par 2 |
||
| + | SEQ-ts SEQ SEQ 3 Par 2 |
||
| + | SEQ2-2opt SEQ SEQ 2 Par 2 |
||
| + | SEQ2-ts SEQ SEQ 3 Par 2 |
||
| + | HC.1.x.10-2opt HC 1.x.10 2 Par 2 |
||
| + | HC.2.6.10-2opt HC 2.6.10 2 Par 2 |
||
| + | HC.2.7.10-2opt HC 2.7.10 2 Par 2 |
||
</pre> |
</pre> |
||
| Line 136: | Line 154: | ||
dev.off() |
dev.off() |
||
} |
} |
||
| ⚫ | |||
| − | |||
| − | ====Things to improve==== |
||
| − | |||
| − | Use a separate text file that contains all the id data on an algorithm (idalgo, topo, schema, ls, type) to reduce the size of the results text files (for the QAP experiments I obtained something like 500MB of text results files, half of which are redundant data) |
||
| ⚫ | |||
| − | result.txt should look like: |
||
| ⚫ | |||
| − | ^^^^^^ ^^^^^^^^ |
||
| − | algorithms.txt should look like: |
||
| ⚫ | |||
| − | ^^^^^ |
||
| − | |||
| − | instances.txt should look like: |
||
| − | instance best-known-solution-value |
||
| − | ^^^^^^^^ |
||
</pre> |
</pre> |
||
Revision as of 22:15, 31 August 2006
Max's scripts
When testing different algorithms on some combinatorial optimization problem, I usually organize the data on my machine according to a same schema. Basically I create the following structure of directories to store sources, problem instances, output data, executables and scripts.
+ main_project_folder | |-+ bin (contains the executables of the algorithms) | |-+ instances (contains the problem instances) | |-+ out (contains the outputs of the trial) | |-+ sh (contains the script to launch the experiments) | |-+ src (contains the sources of the algorithms) | |-+ analysis (contains the R script to boxplot the experimental data)
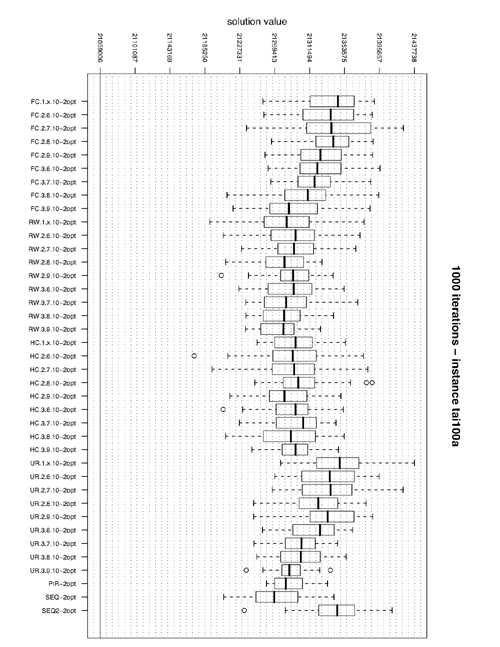
Boxplot of solution values by algorithm
I organize the data in the following way:
in the file optimal_values.txt I put the best-known solution values for the problem instances I'm testing
cat optimal_values.txt instance optimum sko100a 152002 sko100e 149150 tai100a 21059006 tai100b 1185996137
in the files "algorithm.txt" I put the algorithms details
landau:~/Desktop/Parallel-QAP-2cpu/out/analysis/2-opt mmanfrin$ head algorithms.txt idalgo topo schema ls type number_of_cpus PIR-2opt PIR PIR 2 Par 2 PIR-ts PIR PIR 3 Par 2 SEQ-2opt SEQ SEQ 2 Par 2 SEQ-ts SEQ SEQ 3 Par 2 SEQ2-2opt SEQ SEQ 2 Par 2 SEQ2-ts SEQ SEQ 3 Par 2 HC.1.x.10-2opt HC 1.x.10 2 Par 2 HC.2.6.10-2opt HC 2.6.10 2 Par 2 HC.2.7.10-2opt HC 2.7.10 2 Par 2
in the files algorithm_factors_instance-size_cut-time.txt I record the history of the search of the algo for the instance
head sequential_2-opt_100_8000.txt idalgo topo schema ls type cpu_id instance try best time iteration SEQ-2opt SEQ SEQ 2 Seq 0 sko100a 1 155468 0.31 1 SEQ-2opt SEQ SEQ 2 Seq 0 sko100a 1 155390 0.31 1 SEQ-2opt SEQ SEQ 2 Seq 0 sko100a 1 155000 0.31 1 SEQ-2opt SEQ SEQ 2 Seq 0 sko100a 1 154934 0.64 2 SEQ-2opt SEQ SEQ 2 Seq 0 sko100a 1 154608 0.97 3 SEQ-2opt SEQ SEQ 2 Seq 0 sko100a 1 153958 1.27 4 SEQ-2opt SEQ SEQ 2 Seq 0 sko100a 1 153750 1.93 6 SEQ-2opt SEQ SEQ 2 Seq 0 sko100a 1 153720 1.93 6 SEQ-2opt SEQ SEQ 2 Seq 0 sko100a 1 153634 2.57 8
In order to produce boxplots like the one above you can look at the R source used to produce it.
optimal_values<-read.table("optimal_values_100.txt",header=TRUE)
resPIR2OPT<-read.table("parallel_independent_2-opt_100_100.txt",header=TRUE)
resSEQ2OPT<-read.table("sequential_2-opt_100_800.txt",header=TRUE)
resSEQ22OPT<-read.table("sequential2_2-opt_100_100.txt",header=TRUE)
resFC1x102OPT<-read.table("fc.1.x.10_2-opt_100_100.txt",header=TRUE)
resFC26102OPT<-read.table("fc.2.6.10_2-opt_100_100.txt",header=TRUE)
resFC27102OPT<-read.table("fc.2.7.10_2-opt_100_100.txt",header=TRUE)
resFC28102OPT<-read.table("fc.2.8.10_2-opt_100_100.txt",header=TRUE)
resFC29102OPT<-read.table("fc.2.9.10_2-opt_100_100.txt",header=TRUE)
resFC36102OPT<-read.table("fc.3.6.10_2-opt_100_100.txt",header=TRUE)
resFC37102OPT<-read.table("fc.3.7.10_2-opt_100_100.txt",header=TRUE)
resFC38102OPT<-read.table("fc.3.8.10_2-opt_100_100.txt",header=TRUE)
resFC39102OPT<-read.table("fc.3.9.10_2-opt_100_100.txt",header=TRUE)
resHC1x102OPT<-read.table("hc.1.x.10_2-opt_100_100.txt",header=TRUE)
resHC26102OPT<-read.table("hc.2.6.10_2-opt_100_100.txt",header=TRUE)
resHC27102OPT<-read.table("hc.2.7.10_2-opt_100_100.txt",header=TRUE)
resHC28102OPT<-read.table("hc.2.8.10_2-opt_100_100.txt",header=TRUE)
resHC29102OPT<-read.table("hc.2.9.10_2-opt_100_100.txt",header=TRUE)
resHC36102OPT<-read.table("hc.3.6.10_2-opt_100_100.txt",header=TRUE)
resHC37102OPT<-read.table("hc.3.7.10_2-opt_100_100.txt",header=TRUE)
resHC38102OPT<-read.table("hc.3.8.10_2-opt_100_100.txt",header=TRUE)
resHC39102OPT<-read.table("hc.3.9.10_2-opt_100_100.txt",header=TRUE)
resRW1x102OPT<-read.table("rw.1.x.10_2-opt_100_100.txt",header=TRUE)
resRW26102OPT<-read.table("rw.2.6.10_2-opt_100_100.txt",header=TRUE)
resRW27102OPT<-read.table("rw.2.7.10_2-opt_100_100.txt",header=TRUE)
resRW28102OPT<-read.table("rw.2.8.10_2-opt_100_100.txt",header=TRUE)
resRW29102OPT<-read.table("rw.2.9.10_2-opt_100_100.txt",header=TRUE)
resRW36102OPT<-read.table("rw.3.6.10_2-opt_100_100.txt",header=TRUE)
resRW37102OPT<-read.table("rw.3.7.10_2-opt_100_100.txt",header=TRUE)
resRW38102OPT<-read.table("rw.3.8.10_2-opt_100_100.txt",header=TRUE)
resRW39102OPT<-read.table("rw.3.9.10_2-opt_100_100.txt",header=TRUE)
resUR1x102OPT<-read.table("ur.1.x.10_2-opt_100_100.txt",header=TRUE)
resUR26102OPT<-read.table("ur.2.6.10_2-opt_100_100.txt",header=TRUE)
resUR27102OPT<-read.table("ur.2.7.10_2-opt_100_100.txt",header=TRUE)
resUR28102OPT<-read.table("ur.2.8.10_2-opt_100_100.txt",header=TRUE)
resUR29102OPT<-read.table("ur.2.9.10_2-opt_100_100.txt",header=TRUE)
resUR36102OPT<-read.table("ur.3.6.10_2-opt_100_100.txt",header=TRUE)
resUR37102OPT<-read.table("ur.3.7.10_2-opt_100_100.txt",header=TRUE)
resUR38102OPT<-read.table("ur.3.8.10_2-opt_100_100.txt",header=TRUE)
resUR39102OPT<-read.table("ur.3.9.10_2-opt_100_100.txt",header=TRUE)
res<-rbind(resFC1x102OPT,resFC26102OPT,resFC27102OPT,resFC28102OPT,resFC29102OPT,resFC36102OPT,
resFC37102OPT,resFC38102OPT,resFC39102OPT,resRW1x102OPT,resRW26102OPT,resRW27102OPT,resRW28102OPT,
resRW29102OPT,resRW36102OPT,resRW37102OPT,resRW38102OPT,resRW39102OPT,resHC1x102OPT,resHC26102OPT,
resHC27102OPT,resHC28102OPT,resHC29102OPT,resHC36102OPT,resHC37102OPT,resHC38102OPT,resHC39102OPT,
resUR1x102OPT,resUR26102OPT,resUR27102OPT,resUR28102OPT,resUR29102OPT,resUR36102OPT,resUR37102OPT,
resUR38102OPT,resUR39102OPT,resPIR2OPT,resSEQ2OPT,resSEQ22OPT)
linstance<-levels(res$instance)
res.split<-split(1:nrow(res), list(res$instance, res$try, res$idalgo), drop=TRUE)
min.list <- lapply(res.split, function(x){
x[match(min(res$best[x]), res$best[x])]
})
min.vector <- unlist(min.list)
bestalgo<-res[min.vector,]
bestalgo.split <- split(1:nrow(bestalgo), bestalgo$instance, drop=TRUE)
for (i in (1:length(bestalgo.split)))
{
bestalgo.vector <- unlist(bestalgo.split[i])
bestalgo.temp <- bestalgo[bestalgo.vector,]
l<-split(bestalgo.temp$best,bestalgo.temp$idalgo)
epsfile=paste(linstance[i],"_100_nolim.eps",sep="")
postscript(file=epsfile,onefile=TRUE,horizontal=TRUE)
par(mar=c(5,5,5,3),cex.axis=0.7,las=2,mgp=c(4, 1, 0))
title_plot=paste("100 iterations - instance ",linstance[i],sep="")
boxplot(l,xlab="",ylab="solution value",names=c(levels(bestalgo$idalgo)),main=title_plot,
yaxt="n",ylim=c(optimal_values[optimal_values$instance==linstance[i],]$optimum,max(bestalgo.temp$best)))
axis(2, seq(from=optimal_values[optimal_values$instance==linstance[i],]$optimum,to=max(bestalgo.temp$best),length.out=10))
abline(h=optimal_values[optimal_values$instance==linstance[i],]$optimum)
# draw an orizontal line at the y-level of the best-know solution value
grid(nx=0, ny=55,col="gray5")
dev.off()
}