Repository of scripts used by IRIDIA members
From IridiaWiki
The printable version is no longer supported and may have rendering errors. Please update your browser bookmarks and please use the default browser print function instead.
Max's scripts
When testing different algorithms on some combinatorial optimization problem, I usually organize the data on my machine according to a same schema. Basically I create the following structure of directories to store sources, problem instances, output data, executables and scripts.
+ main_project_folder | |-+ bin (contains the executables of the algorithms) | |-+ instances (contains the problem instances) | |-+ out (contains the outputs of the trial) | | | |-+ algorithm1 | | | | | |-+ instance1 | | | | | |-+ instance2 | | | |-+ algorithm2 | | | | | |-+ instance1 | | | | | |-+ instance2 | |-+ sh (contains the script to launch the experiments) | |-+ src (contains the sources of the algorithms) | |-+ algorithm1 | | | |-+ algorithm2 | |-+ analysis (contains the R script to boxplot the experimental data)
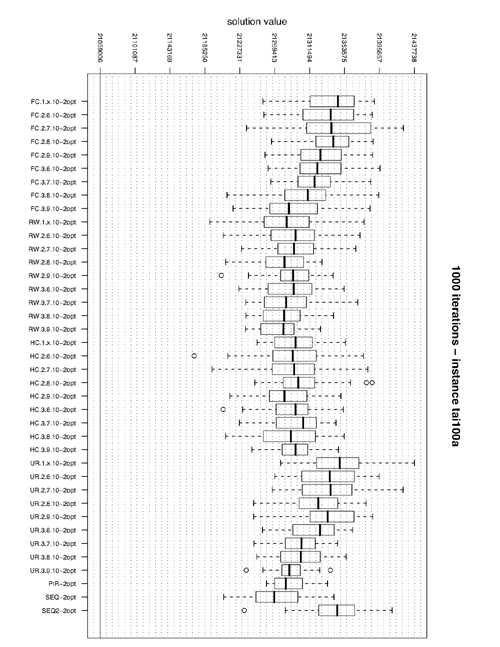
Boxplot of solution values by algorithm
I organize the data in the following way:
in the file optimal_values.txt I put the best-known solution values for the problem instances I'm testing
instance optimum sko100a 152002 sko100e 149150 tai100a 21059006 tai100b 1185996137
in the files algorithms.txt I put the algorithms details
idalgo topo schema ls type number_of_cpus PIR-2opt PIR PIR 2 Par 2 PIR-ts PIR PIR 3 Par 2 SEQ-2opt SEQ SEQ 2 Par 2 SEQ-ts SEQ SEQ 3 Par 2 SEQ2-2opt SEQ SEQ 2 Par 2 SEQ2-ts SEQ SEQ 3 Par 2 HC.1.x.10-2opt HC 1.x.10 2 Par 2 HC.2.6.10-2opt HC 2.6.10 2 Par 2 HC.2.7.10-2opt HC 2.7.10 2 Par 2
in the files algorithm_factors_instance-size_cut-time.txt I record the history of the search of the algo for the instance
idalgo cpu_id instance try best time iteration HC.1.x.10-2opt 0 sko100a 1 155254 0.35 1 HC.1.x.10-2opt 0 sko100a 1 154162 0.35 1 HC.1.x.10-2opt 0 sko100a 1 154050 0.35 1 HC.1.x.10-2opt 0 sko100a 1 153684 2.69 8 HC.1.x.10-2opt 0 sko100a 1 153508 3.24 10 HC.1.x.10-2opt 0 sko100a 1 153396 3.24 10 HC.1.x.10-2opt 0 sko100a 1 153344 3.24 10 HC.1.x.10-2opt 0 sko100a 1 153092 3.52 11 HC.1.x.10-2opt 0 sko100a 1 153062 3.75 12
In order to produce boxplots like the one above you can look at the R source used to produce it.
optimal_values<-read.table("optimal_values.txt",header=TRUE)
resPIR2OPT<-read.table("parallel_independent_2-opt_100_1000.txt",header=TRUE)
resSEQ2OPT<-read.table("sequential_2-opt_100_2000.txt",header=TRUE)
resSEQ22OPT<-read.table("sequential2_2-opt_100_1000.txt",header=TRUE)
resHC1x102OPT<-read.table("hc.1.x.10_2-opt_100_1000.txt",header=TRUE)
resHC26102OPT<-read.table("hc.2.6.10_2-opt_100_1000.txt",header=TRUE)
resHC27102OPT<-read.table("hc.2.7.10_2-opt_100_1000.txt",header=TRUE)
resHC28102OPT<-read.table("hc.2.8.10_2-opt_100_1000.txt",header=TRUE)
resHC29102OPT<-read.table("hc.2.9.10_2-opt_100_1000.txt",header=TRUE)
resHC36102OPT<-read.table("hc.3.6.10_2-opt_100_1000.txt",header=TRUE)
resHC37102OPT<-read.table("hc.3.7.10_2-opt_100_1000.txt",header=TRUE)
resHC38102OPT<-read.table("hc.3.8.10_2-opt_100_1000.txt",header=TRUE)
resHC39102OPT<-read.table("hc.3.9.10_2-opt_100_1000.txt",header=TRUE)
resRW1x102OPT<-read.table("rw.1.x.10_2-opt_100_1000.txt",header=TRUE)
resRW26102OPT<-read.table("rw.2.6.10_2-opt_100_1000.txt",header=TRUE)
resRW27102OPT<-read.table("rw.2.7.10_2-opt_100_1000.txt",header=TRUE)
resRW28102OPT<-read.table("rw.2.8.10_2-opt_100_1000.txt",header=TRUE)
resRW29102OPT<-read.table("rw.2.9.10_2-opt_100_1000.txt",header=TRUE)
resRW36102OPT<-read.table("rw.3.6.10_2-opt_100_1000.txt",header=TRUE)
resRW37102OPT<-read.table("rw.3.7.10_2-opt_100_1000.txt",header=TRUE)
resRW38102OPT<-read.table("rw.3.8.10_2-opt_100_1000.txt",header=TRUE)
resRW39102OPT<-read.table("rw.3.9.10_2-opt_100_1000.txt",header=TRUE)
res<-rbind(resRW1x102OPT,resRW26102OPT,resRW27102OPT,resRW28102OPT,resRW29102OPT,
resRW36102OPT,resRW37102OPT,resRW38102OPT,resRW39102OPT,resHC1x102OPT,resHC26102OPT,
resHC27102OPT,resHC28102OPT,resHC29102OPT,resHC36102OPT,resHC37102OPT,resHC38102OPT,
resHC39102OPT,resPIR2OPT,resSEQ2OPT,resSEQ22OPT)
linstance<-levels(res$instance)
res.split<-split(1:nrow(res), list(res$instance, res$try, res$idalgo), drop=TRUE)
min.list <- lapply(res.split, function(x){
x[match(min(res$best[x]), res$best[x])]
})
# matches return the first among all the values with min best!!!
# so is not the one with minimal time
min.vector <- unlist(min.list)
bestalgo<-res[min.vector,]
bestalgo.split <- split(1:nrow(bestalgo), bestalgo$instance, drop=TRUE)
for (i in (1:length(bestalgo.split)))
{
bestalgo.vector <- unlist(bestalgo.split[i])
bestalgo.temp <- bestalgo[bestalgo.vector,]
l<-split(as.double(bestalgo.temp$best),bestalgo.temp$idalgo)
epsfile=paste(linstance[i],"_1000.eps",sep="")
postscript(file=epsfile,onefile=TRUE,horizontal=TRUE)
par(mar=c(5,5,5,3),cex.axis=0.7,las=2,mgp=c(4, 1, 0))
title_plot=paste("2 CPU - 1000 iterations - instance ",linstance[i],sep="")
boxplot(l,xlab="",ylab="solution value",names=c(levels(bestalgo$idalgo)),main=title_plot,
yaxt="n",ylim=c(min(min(bestalgo.temp$best),optimal_values[optimal_values$instance==linstance[i],]$optimum),
max(bestalgo.temp$best)))
axis(2, seq(from=min(min(bestalgo.temp$best),optimal_values[optimal_values$instance==linstance[i],]$optimum),
to=max(bestalgo.temp$best),length.out=10))
abline(h=optimal_values[optimal_values$instance==linstance[i],]$optimum)
grid(nx=0, ny=55,col="gray5")
dev.off()
}